Transcriptomics
COMMON SUPPORT PLATFORMS
Equipment
The massive sequencing system (Next Generation Sequencing or NGS) has a wide range of applications, increasingly numerous, in all studies or research projects that involve nucleic acid analysis. Currently, this platform is equipped with an Illumina NextSeq 2000 System, which offers all the advantages of this leading technology and with which promising results are being achieved in recent years.
Description
The technology used by the Illumina NextSeq2000 is based on sequencing by synthesis (SBS), where tagged nucleotides are tracked simultaneously with the copying of the DNA strand. 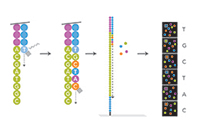 This technology allows millions of fragments to be sequenced massively and in parallel, improving the speed and accuracy of sequencing while reducing its cost. The Illumina NextSeq 2000 Sequencing System has been provided with a novel super-resolution optical system that produces high-precision image data with higher resolution and sensitivity than more traditional Illumina systems. This technology also provides greater sequencing flexibility, scalable to different production experimental needs, and adaptable to both conventional and emerging applications. (Video https://youtu.be/fCd6B5HRaZ8) The flexibility of the technology makes the platform adaptable to a wide range of applications and applicable to projects with diverse sequencing objectives: Transcriptome sequencing The Nextseq 2000 System offers great flexibility for RNA sequencing studies, which require different sequencing depths and read lengths depending on study complexity, size, and goals: • Total RNA sequencing. Analyze the coding regions and multiple forms of non-coding RNA to get a complete picture of the transcriptome. • mRNA sequencing. Detect and quantify new and known transcripts for complete and accurate expression profiles. • Gene expression profiles. It allows multiple RNA libraries to be sequenced together in a single sequencing run, obtaining a rapid picture of the most expressed genes with only a few reads per sample. • Targeted RNA sequencing. Select and sequence transcripts of specific interest for gene expression profiling studies. • Sequencing of small RNA (RNAs) and MicroRNA (miRNA). It allows querying thousands of RNA and microRNA sequences with high sensitivity. Sequencing of the human exome and genome of microorganisms • Sequencing of small genomes. It sequences the entire genome of a bacterium, virus, or other microbes. • Sequencing of the human exome. Captures and sequences the protein coding region of the genome, a cost-effective alternative to whole genome sequencing. Targeted panel sequencing The gene targeting and gene targeting panels sequence only genomic or transcriptomic regions of interest. They are useful tools to analyze specific mutations, mutational load, microsatellite instability, HLA genotyping, expression of the TCR repertoire or gene fusion profiles…. Applications of targeted panel sequencing include: • Personalized genomic sequencing. Selection and sequencing of specific regions of the genome that have a specific research interest: for example, specific biological pathways or follow-up experiments from genome-wide association studies or whole genome sequencing. • Pre-designed panels for targeted gene sequencing. They contain fixed genomic regions or genes associated with a disease, phenotype, or biological pathway. They are available for research of various diseases like cancer, hereditary disorders, rare disorders, cardiac disorders, neurological disorders, immune diseases, etc. Other possible applications -Free circulating DNA sequencing and liquid biopsy analysis -Chromatin analysis (ChIP-Seq, ATAC-Seq) -DNA methylation sequencing -Sequencing of 16S ribosomal RNA (rRNA) and internal transcribed spacer (ITS) -Metagenomic profiles (metagenomics, metatranscriptomics) -Single cell DNA and RNA sequencing
This technology allows millions of fragments to be sequenced massively and in parallel, improving the speed and accuracy of sequencing while reducing its cost. The Illumina NextSeq 2000 Sequencing System has been provided with a novel super-resolution optical system that produces high-precision image data with higher resolution and sensitivity than more traditional Illumina systems. This technology also provides greater sequencing flexibility, scalable to different production experimental needs, and adaptable to both conventional and emerging applications. (Video https://youtu.be/fCd6B5HRaZ8) The flexibility of the technology makes the platform adaptable to a wide range of applications and applicable to projects with diverse sequencing objectives: Transcriptome sequencing The Nextseq 2000 System offers great flexibility for RNA sequencing studies, which require different sequencing depths and read lengths depending on study complexity, size, and goals: • Total RNA sequencing. Analyze the coding regions and multiple forms of non-coding RNA to get a complete picture of the transcriptome. • mRNA sequencing. Detect and quantify new and known transcripts for complete and accurate expression profiles. • Gene expression profiles. It allows multiple RNA libraries to be sequenced together in a single sequencing run, obtaining a rapid picture of the most expressed genes with only a few reads per sample. • Targeted RNA sequencing. Select and sequence transcripts of specific interest for gene expression profiling studies. • Sequencing of small RNA (RNAs) and MicroRNA (miRNA). It allows querying thousands of RNA and microRNA sequences with high sensitivity. Sequencing of the human exome and genome of microorganisms • Sequencing of small genomes. It sequences the entire genome of a bacterium, virus, or other microbes. • Sequencing of the human exome. Captures and sequences the protein coding region of the genome, a cost-effective alternative to whole genome sequencing. Targeted panel sequencing The gene targeting and gene targeting panels sequence only genomic or transcriptomic regions of interest. They are useful tools to analyze specific mutations, mutational load, microsatellite instability, HLA genotyping, expression of the TCR repertoire or gene fusion profiles…. Applications of targeted panel sequencing include: • Personalized genomic sequencing. Selection and sequencing of specific regions of the genome that have a specific research interest: for example, specific biological pathways or follow-up experiments from genome-wide association studies or whole genome sequencing. • Pre-designed panels for targeted gene sequencing. They contain fixed genomic regions or genes associated with a disease, phenotype, or biological pathway. They are available for research of various diseases like cancer, hereditary disorders, rare disorders, cardiac disorders, neurological disorders, immune diseases, etc. Other possible applications -Free circulating DNA sequencing and liquid biopsy analysis -Chromatin analysis (ChIP-Seq, ATAC-Seq) -DNA methylation sequencing -Sequencing of 16S ribosomal RNA (rRNA) and internal transcribed spacer (ITS) -Metagenomic profiles (metagenomics, metatranscriptomics) -Single cell DNA and RNA sequencing
Services
Our service includes all the stages of support that your project needs, from consultancy and experimental advice, to technical support for the development of sequencing protocols, material management, sample analysis and the corresponding analysis of bioinformatic data for its publication. Experimental advice Our sequencing platform provides experimental advice for a wide variety of research projects that require: • Total RNA sequencing • mRNA sequencing • Sequencing of small RNAs and miRNAs • Gene expression profiles • Complete sequencing of small genomes •Human whole exome sequencing (based on enrichment technology) • Sequencing of specific gene panels (based on PCR amplicon or enrichment technology) • Sequencing of ctDNA, Chip-Seq, ATAQ-seq and any other genomic application, with prior evaluation of the request. Technical support We offer technical support for the complete development of the sequencing protocol: • Experimental design of sequencing projects • Extraction of nucleic acids from different types of samples: blood, saliva, bone or tissue embedded in formalin fixed paraffin (FFPE). • Quality control and integrity of the samples • Preparation and quality control of the NGS libraries • Sequencing of NGS libraries Materials management We provide the management and ordering of the necessary reagents for the preparation of the NGS libraries, flow cells and cartridges for the required sequencing application. Bioinformatics data analysis We deliver the results in sequencing file formats containing the raw data and offer specialized processing and analysis of NGS data: quality control, de-multiplexing, alignment and post-analysis.
Fees
VAT not included



